August 26, 2020
New insights into the SARS-CoV-2 spike protein
The "gate opener" on the virus surface is unexpectedly mobile
The search for a suitable vaccine is a central research area in the fight against the coronavirus SARS-CoV-2. An important step towards this is to understand how the virus penetrates the cells. The protein responsible for this is the SARS-CoV-2 spike (S) protein on the virus surface which allows the virus to enter the host cell.
To ease the development of a protective vaccine, FIAS scientists Gerhard Hummer and Roberto Covino use advanced computer simulations to identify weak points on the surface of the S protein that could be attacked by antibodies. In collaboration with scientists from the Max Planck Institute for Biophysics, EMBL, the Paul Ehrlich Institute (PEI), and other research institutions, they have comprehensively analyzed and modeled the spike protein. In the online edition of Science from August 18, 2020, they report on their research and the finding that the spike protein is much more mobile than first assumed.
Due to the collaboration of various expertise, the protein could be analyzed in its natural environment with imaging techniques as well as with computer simulations. To determine the dynamics and structure of the spike, the biophysicists and Gerhard Hummer and Roberto Covino, together with their colleagues from the MPI for Biophysics, applied computer model simulations from membrane dynamics to the protein. At the same time, researchers at the PEI extracted SARS-CoV-2 particles from the supernatant of infected cells, which were then accurately mapped in situ (on the virus) by EMBL and MPI scientists using cryo-electron microscopy. These 3-dimensional images of the protein on the surface of the virus were then integrated with the theoretical model of the Frankfurt group to elucidate the structure and dynamics of the protein in detail.
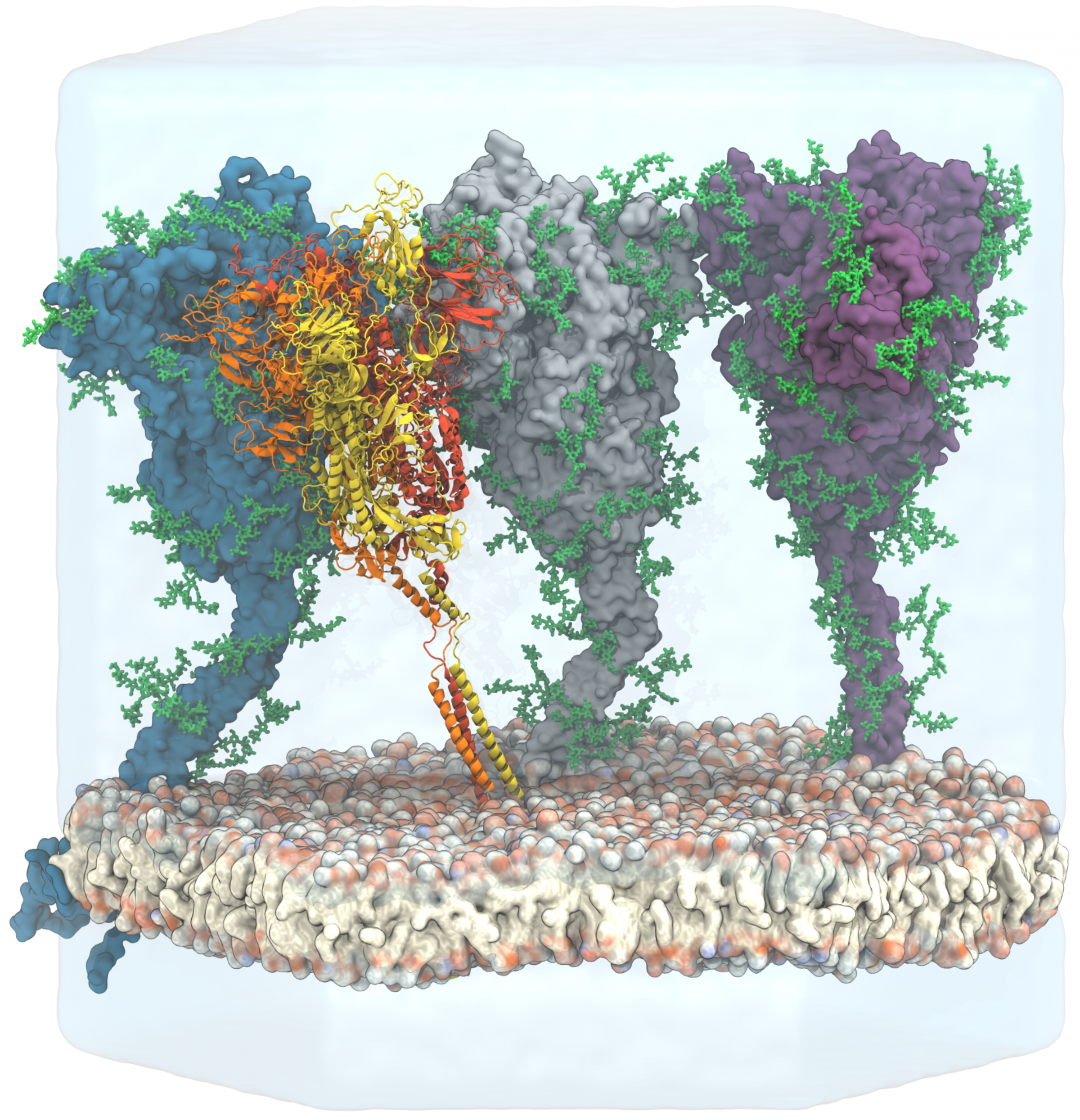
From the models and images, the researchers were able to conclude that the protein is much more flexible than previously assumed. This property enables the virus to dock to human cells like an anchor hook and enter them. Also, the interdisciplinary collaboration has discovered sugar-like deposits on the protein. These so-called glycans hide the protein structure from the immune system and make it difficult for the antibodies to dock to the protein. Such information is important for assessing the effectiveness of vaccines or antibody therapies.
Further Information:
Publication: https://science.sciencemag.org/content/early/2020/08/17/science.abd5223
Press release by the Paul-Ehrlich Institute: https://www.pei.de/EN/newsroom/press-releases/year/2020/14-new-findings-spike-protein-sars-cov-2.html
Picture: Sikora et.al; Map of SARS-CoV-2 spike epitopes not shielded by glycans Licence: Creative Commons CC-BY-NC-ND 4.0